A difficult task: Phylogenetic analysis of SARS-CoV-2 data
-
author:
Isabel Lacurie
- place:
- date: 13.08.2020
Computer scientists from HITS and KIT review the challenges for phylogenetic methods in analyzing the evolution of the corona virus: Large number of sequences, low number of mutations.
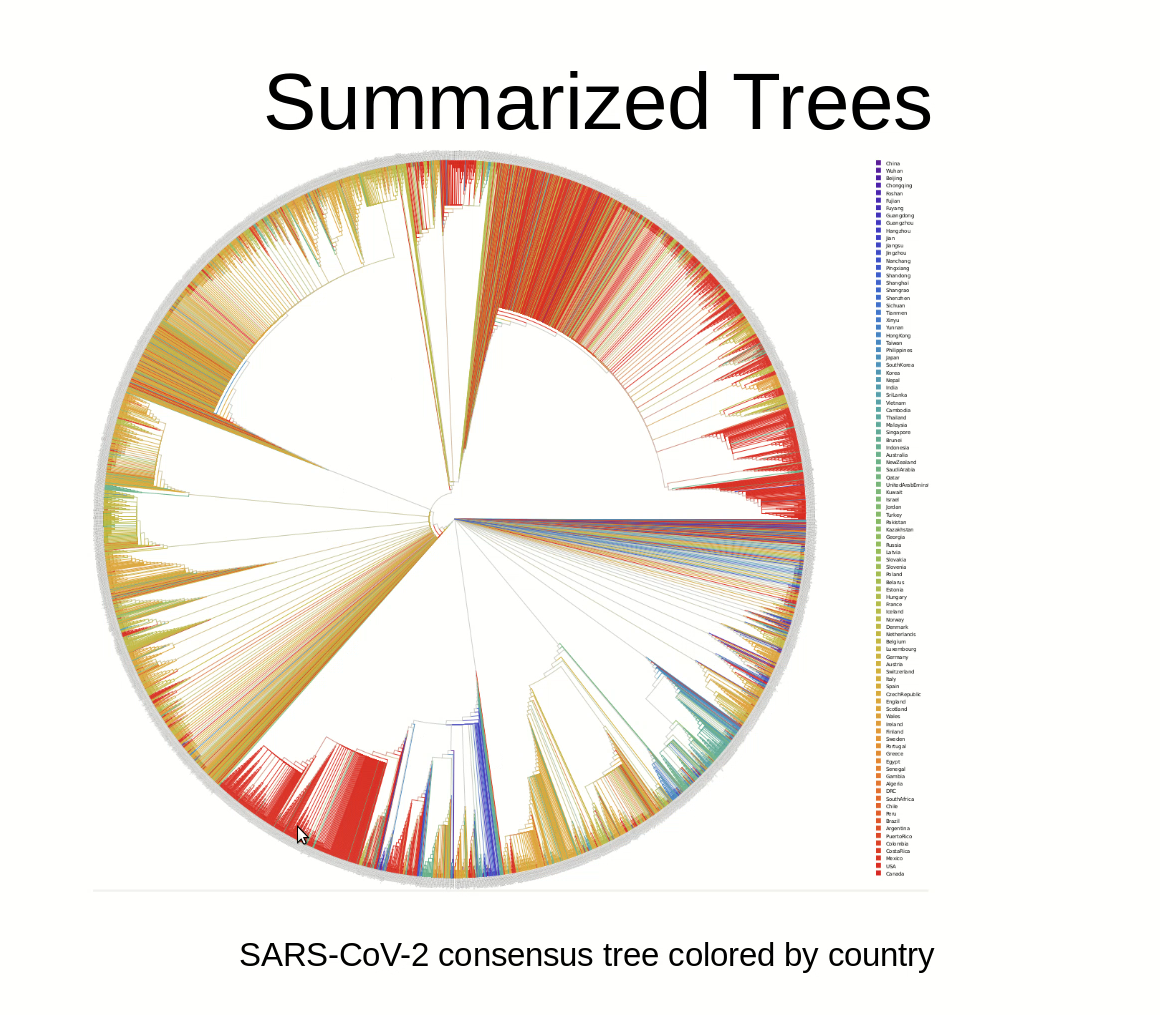
Many studies covering some aspects of SARS-CoV-2 data analyses are being published on a daily basis, including a regularly updated phylogeny. In a preprint paper, researchers from the Computational Molecular Evolution group (CME) at the Heidelberg Institute for Theoretical Studies (HITS) and the Institute for Theoretical Informatics at the Karlsruhe Institute of Technology (KIT) have analyzed all available virus sequences available at the beginning of May. They found it difficult to infer a reliable phylogeny on these data due to the large number of sequences in conjunction with the low number of mutations. “The data contain an extremely weak signal”, says CME group leader Alexandros Stamatakis.”
They also found that it is not possible to reliably root the inferred phylogeny either using the bat and pangolin or by applying novel computational methods on the human virus phylogeny. Even an automatic classification of the current sequences was not possible, as the sequences are too closely related.
The researchers conclude that results of phylogenetic analyses on SARS-CoV-2 data should be considered and interpreted with extreme caution. “Do not draw conclusions from just a single tree”, says Alexandros Stamatakis.
The paper is available on the bioRxiv preprint server:
Phylogenetic analysis of SARS-CoV-2 data is difficult. Benoit Morel, Pierre Barbera, Lucas Czech, Ben Bettisworth, Lukas Hübner, Sarah Lutteropp, Dora Serdari, Evangelia-Georgia Kostaki, Ioannis Mamais, Alexey M Kozlov, Pavlos Pavlidis, Dimitrios Paraskevis, Alexandros Stamatakis: https://www.biorxiv.org/content/10.1101/2020.08.05.239046v1